Accelerating Medical Applications with Xilinx Vitis AI
Objectives
Medical AI is becoming an essential part of modern healthcare. It increases detection accuracy, lowers cost, achieves earlier disease prediction, and largely uses non-invasive means. In this respect, imaging equipment plays a critical role. With the growing applications in this field AI innovations and algorithms associated with medical image processing are evolving rapidly. Hardware implementation of these algorithms offers much better efficiency and lower latency. Xilinx’s full-stack deep learning SDK, Vitis AI, along with highly adaptive Xilinx’s AI platforms, enables medical equipment manufacturers and developers with rapid prototyping of these highly evolving algorithms, minimizing time-to-market and cost. The reconfigurability of Xilinx platforms provides longer life utility of their products as well.
Because of the inherent characteristics of medical images, (higher resolution, largely electronic and photonic 3D format, etc.), these images require significant preprocessing before feeding them into an AI inference engine. These preprocessing algorithms require that several complex operations be performed on each pixel in each image, resulting in a large number of operations per second. Implementing these preprocessing algorithms in Xilinx platforms along with the Xilinx’s AI inference engine (DPUs) provides near real-time detection, which can be essential in medical care. Xilinx’s Vitis AI provides an end-to-end development platform for these whole application acceleration solutions.
In a set of two articles, we demonstrate how a medical application developer can take a medical image dataset and develop and evaluate an end-to-end AI accelerated application using Vitis AI, without writing any lower level RTL code. In this first article, we take the open-source Skin Lesions dataset[1], preprocess the images, create a machine learning network, and train it on an x86 platform. Then we use the Xilinx’s Vitis AI toolset to quantize, compile and evaluate the model in one of the Xilinx’s AI inference platforms, the ZCU102 board. In the second article we are going to create an end-to-end solution where the image preprocessing IP, will perform real-time contrast adjustment, brightness adjustment, and inverting and pseudo color operations before feeding them into the Xilinx AI acceleration IPs.
Introduction
The development of an accelerated deep learning application is a two-phase process. Phase 1 is the model creation and training and Phase 2 is the model deployment for inferencing. During the training phase, we design a deep neural network for a specific task and train it with a large dataset; in our case the task is to classify the skin lesion images. During the inference phase, we compile and deploy the trained model with the application logic targeting a specific AI platform, in our case, it’s the Xilinx’s ZCU102 platform. During inference the application logic feeds new data/images to the model for classification. For our skin lesion detection application, these phases are described in more detail below.
Phase 1: Model Creation and Training
1.0 Design Considerations
1.1 Transfer Learning consideration
Training a deep neural network requires a large dataset. Collecting a large number of medical images is always a challenge. Our skin lesion dataset[1] only has 10k images, which is insufficient for training a deep network from scratch. To overcome this challenge, we use a Transfer Learning technique. Transfer Learning can be achieved in two ways, fine-tuning a network pre-trained on general images or fine-tuning a network pre-trained on medical images for a different target organ or task. For ease of availability, we chose to use an Inception v3 network pre-trained with ImageNet[2], a dataset of over 14 million general images
1.2 Dataset size consideration
The size of the dataset is a key parameter in deciding on the level of Transfer Learning. Even with some augmentation, our skin dataset is small. The number of parameters in the Inception v3 network is very large, roughly 22 million, so full adaption may result in overfitting. Thus, we chose to proceed with partial adaption.
1.3 Vitis AI Considerations
When creating a deep neural network or choosing one for Transfer Learning targeting Xilinx’s AI platforms for deployment, we need to make sure all layers, the size of the kernels, and the activation functions in the network are supported by the Xilinx Vitis AI stack. Our selected Inception v3 for Transfer Learning is fully supported by the Vitis AI stack. Also, in our network modifications both Dense and Softmax layers are supported (more on this coming later). The kernel size and the input/output dimensions of these newly added layers are within the supported range as well. For more details on the supporting layer, kernel size, etc. please refer to Xilinx’s DPU IP Product Guide PG338.[3]
2.0 Data Preparation: Image-preprocessing
Supervised training of neural networks for automated diagnosis of pigmented skin lesions requires homogenous, clean and labeled datasets of images. The skin lesion dataset we use is already preprocessed and released as a training set for academic machine learning purposes and is publicly available through the ISIC archive. The images are collected from different populations, by different medical persons at various times with many different dermatoscopes. Given this diversity, various acquisition and cleaning methods are applied during the preprocessing flow. For example, the specimen (a) in Figure 1 below is an original image taken by a dermatoscope. This image has a black border at the bottom left edge. It also has low luminance, yellow hue and the lesion is off-center. The image (b) at the right is the processed clean image.
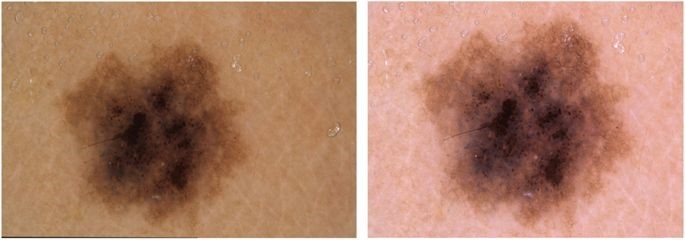
Data cleaning is a very important step for convergence during the training flow. Similarly, clean homogeneous specimens are equally important for better detection accuracy during inference. This preprocessing requires significant computations. In the follow-on second article we demonstrate that by implementing these preprocessing algorithms in the hardware in Xilinx’s platforms, along with Xilinx’s AI inference engine (DPUs), customers can achieve near real-time detection speed.
3.0 Network Design, Training on GPU and Evaluation
The objective of our detection application is to classify skin lesion specimens into seven classes. These seven classes are: nv (Melanocytic nevi), mel (Malignant neoplasm), bkl (Benigh Keratosis), bcc (Basal cell Carcinoma), akiec (Actinic Keratoses), vasc (Vascular Skin) and df (Dermatofibroma).
3.1 Creating a Balanced dataset
Analyzing the skin lesion dataset, we see it consists of 10015 images from these seven classes. Class nv has 6705 images, mel has 1113, bkl has 1099, bcc has 514, akiec has 327, vasc has 142 and df has 115. Clearly this dataset is not balanced.
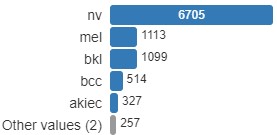
Training the network with this dataset as-is would create a model which is heavily skewed by a single majority class, nv. To prevent this, we use an offline data augmentation technique to reduce the class imbalance. To create augmented images we use the Keras ImageDataGenerator class as follows,
# Data generator
Datagen=ImageDataBenerator(width_shift_range=0.1, height_shift_range=0.1, rotation_range=180, zoom_range=0.1, horizontal_flip=True, vertical_flip=True, fill_mode='nearest')
We also split the dataset into training and validation sets at a factor of 10:3. For details refer to the Jupyter notebook dataaug.ipynb located at the GitHub link in the References section at the end of this article[10].
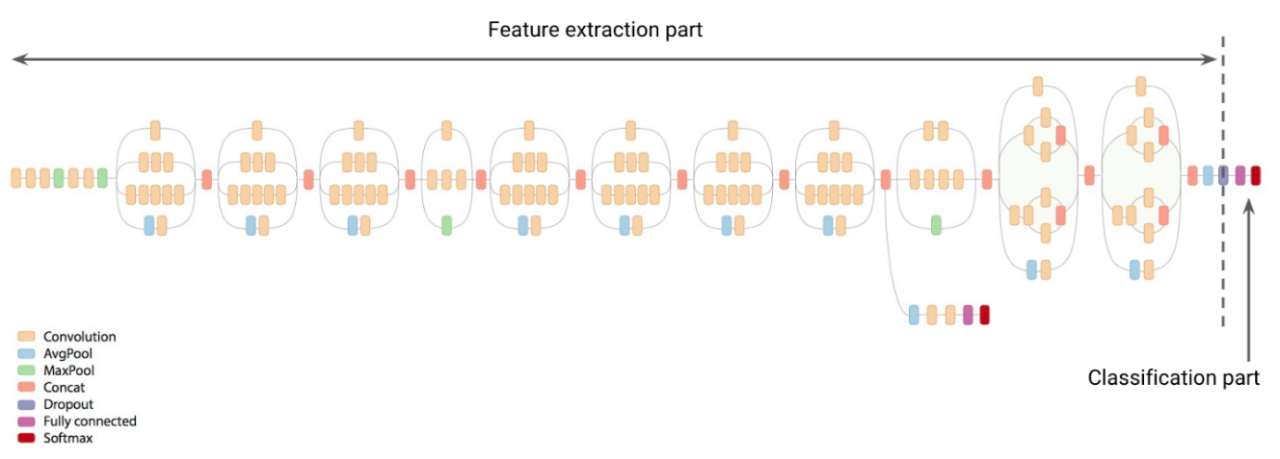
3.2 Network Modification
The publicly available Inception v3 network is designed to classify object images into 1001 output classes. To meet our application requirement, we augment the network to support classification between 7 output classes. First the classification layer of the Inception v3 is replaced with a flattened layer. Than two fully connected layers are added right after the flatten layer. Finally, we add a prediction layer (also called Softmax layer) with 7 outputs, one for each category of skin lesion disease. Here is the Keras code snippet where we create the modified network. For details refer to the Jupyter rnotebook, training.ipynb located at the GitHub link.
# Create an inception_v3 model along with the imagenet pre-trained weights
model=keras.applications.inception_v3.InceptionV3(include_top=False, weights='imagenet', input_shape=(224, 224, 3))
# Taking the output of the inception_v3 just before last layer
x = iv3_model.layers[-1].output
# flattening the outputs of the last conv layer
flatten = Flatten()(x)
# adding two fully connected layers. Meeting DPU requirement, keeping output/input ratio at @ 1/6
dense1 = Dense(2048, activation= 'relu')(flatten)
dense3= Dense(128, activation= 'relu')(dense1)
# adding the prediction layer with 'softmax'
predictions = Dense(7, activation='softmax')(dense3)
# Create the new model with the new outputs
model = Model(inputs=iv3_model.input, outputs=predictions)
3.3 Training Flow
For training, we freeze the pre-trained weights of all the convolutional layers of the Inception V3. We also randomly initialize the weights of the newly added fully connected layers. Since fine-tuning, we choose to use a small learning rate during back-propagation runs. The Categorical Cross-entropy algorithm is used for loss calculation while Adam is used to update the weights of the network. Here is the code snippet that we use for training flow. For details refer to the Jupyter notebook, training.ipynb, also located at the GitHub link.
# Freeze the weights of the layers that we aren't training (training the last 23)
for layer in model.layers[:-3]:
layer.trainable = False
# Compile and prepare the network for training
model.compile(Adam(lr=0.001), loss='categorical_crossentropy', metrics=[categorical_accuracy, top_2_accuracy, top_3_accuracy])
# Declare a checkpoint to save the best trained model
checkpoint = ModelCheckpoint(filepath, monitor='val_categorical_accuracy', verbose=1,
save_best_only=True, mode='max')
# Declare how the training flow reduce the learning rate as the learning platos
reduce_lr = ReduceLROnPlateau(monitor='val_categorical_accuracy', factor=0.2, patience=2,
verbose=1, mode='max', min_lr=0.000001)
callbacks_list = [checkpoint, reduce_lr]
# Train the model for 20 epochs
history = model.fit_generator(train_batches,
steps_per_epoch=train_steps,
class_weight=class_weights,
validation_data=valid_batches,
validation_steps=val_steps,
epochs=20,
verbose=1,
callbacks=callbacks_list)
The training process results in optimal weights and achieves a good classification accuracy of 72.6% and top-3 accuracy of 93.7%.
val_cat_acc: 0.7265469035256409
val_top_2_acc: 0.862275450618681
val_top_3_acc: 0.9371257461533099
Since we use a Keras framework for the development of the classifier model, the output of the trained weight file is in HDF5 or .h5 format. The Vitis AI quantizer tool accepts .pb format, also known as frozen graph format. We use the following code snippet to convert .h5 file and the network into .pb format. For details refer to training.ipnb.
from keras import backend as K
frozen_graph = freeze_session(K.get_session(),output_names=[out.op.name for out in model.outputs])
tf.train.write_graph(frozen_graph, ".", "model/model_incv3_2_6.pb", as_text=False)
Phase 2: Port and Evaluate the Model on Xilinx’s AI platforms
The key advantages of deploying machine learning models on Xilinx’s AI platforms are better inference latency and smaller power & memory footprint. We achieve these by quantizing the 32-bit floating point weights of the trained model into lower precision. Also, Xilinx’s deep learning processor unit (DPU) is a programmable engine that executes a highly optimized instruction set. Hence, we need to compile the modified Inception v3 network into instruction sets that can be executed by the DPU. Xilinx’s Vitis AI toolchain comes with all the necessary tools to quantize and compile our trained model.
1.0 Vitis AI tools Bring Up
The Vitis AI software is available as a Docker image and is available for download from Docker hub[6]. To install Docker we follow the instructions available at www.docs.Docker.com [7]. We clone the Vitis AI 1.0 GitHub repository and pull Xilinx/vitis-aitools-1.0.0-cpu Docker image from Docker hub. Step by step instructions are available in the Vitis AI guide UG1414 [5]. Here are the command snippets:
# pull the vitis-ai tools image from Docker hub
$ Docker pull xilinx/vitis-ai:tools-1.0.0-cpu
#Clone the Vitis AI repository
$ git clone https://github.com/xilinx/vitis-ai
# Launch the Docker container
$ cd Vitis-AI
$ ./Docker_run.sh xilinx/vitis-ai:tools-1.0.0-cpu
# Once inside the container at /workspace, activate the vitis-ai-tensorflow conda environment
$ conda activate vitis-ai-tensorflow
2.0 Quantization of the Weights
Currently Xilinx’s AI platforms support 8 bit precision. To quantize the trained weights we took advantage of the via_q_tensorflow application that comes pre-installed in the Vitis AI tools Docker. We call the via_q_tensorflow with the parameters shown in the table below. For additional options and parameters refer to UG1414.
# Quantize the trained weights of skinl model
$ vai_q_tensorflow quantize \
--input_frozen_graph ${TF_NETWORK_PATH}/${FROZEN_MODEL} \
--input_fn ${INPUT_FN} \
--input_nodes ${INPUT_NODES} \
--output_nodes ${OUTPUT_NODES}\
--input_shapes ?,224,224,3 \
--calib_iter 10 \
--method 1 \
--gpu 0 \
--output_dir ${TF_NETWORK_PATH}/qoutput
Name |
Type |
Description |
--input_frozen_graph |
String |
Frozen inference GraphDef file; the model/model_incv3_2_6.pb we created during Phase 1 |
--input_nodes |
String |
A list of input node names of the frozen graph |
--output_nodes |
String |
A list of output nodes of the frozen graph |
--input_shapes |
String |
A list of 4d shapes of all input_nodes. |
--input_fn |
String |
This function feeds the input calibration images to the graph. The function format is module_name.input_fn_name. |
--method |
Int32 |
The method for quantization. We use 0: Non-overflow method |
--calib_iter |
Int32 |
The iterations of calibration. Total number of images for calibration = calib_iter * batch_size. |
--output_dir |
string |
The directory in which to save the quantization results. |
We gather the input_node name, the output_node name and input_shape by executing the vitis_ai_tensorflow application with ‘inspect’ option as shown below,
$ vai_q_tensorflow inspect
--input_frozen_graph=model_incv3_2_6.pb
Op types used: 912 Const, 350 Identity, 87 Relu, 86 Conv2D, 86 Fill, 86 FusedBatchNorm, 12 ConcatV2, 9 AvgPool, 4 MaxPool, 3 BiasAdd, 3 MatMul, 1 Pack, 1 Placeholder, 1 Prod, 1 Reshape, 1 Shape, 1 Softmax, 1 StridedSlice
Found 1 possible inputs: (name=input_1, type=float(1), shape=[?,224,224,3])
Found 1 possible outputs: (name=dense_3/Softmax, op=Softmax)
The calibration image set is a subset of the training/validation dataset or actual application images. We copied 300 calibration images during the ‘data augmentation’ step in Phase-1. For details refer to the Jupyter notebook dataaug.ipynb.
For input_fn we modified the input_fn.py that comes with the Vitis AI examples in the repository. We modify the calib_input(..) function in the input_fn.py. It reads the calibration images batch by batch and returns a numpy dictionary of {input_node_name, numpy.Array_images}
For convenience, we create a bash script dnnq_skinl.sh with the quantization function call. Once we have all necessary parameters we update the script and execute it within the Docker container under conda environment vitis-ai-tensorflow
$ ./dnnq_skinl.sh
Once the quantization is done, the summary displays as shown below:
INFO: Calibration Done.
INFO: Generating Deploy Model...
INFO: Deploy Model Generated.
********************* Quantization Summary *********************
INFO: Output:
quantize_eval_model: qoutput/quantize_eval_model.pb
deploy_model: qoutput/deploy_model.pb
During quantization two files are generated in the output directory named qoutput/. The deploy_model.pb file contains the quantized trained weights. We use this file in the next section during network compilation. The quantize_eval_model.pd file can be used for model evaluation which we will cover in a future article.
3.0 Compilation of the Network Graph
The Vitis AI tools Docker comes with Vitis AI VAI_C, a domain-specific compiler. It efficiently maps the network model into a highly optimized instruction sequence for the Xilinx’s Deep learning Processor Unit (DPU). We call the via_c_tensorflow with the parameters shown in the table below. For additional options and parameters refer to UG1414.
# Compile the skinl network
$ vai_c_tensorflow --frozen_pb qoutput/deploy_model.pb \
--arch {ARCH} \
--output_dir coutput/output_ZCU102/ \
--net_name skinl
Parameters |
Description |
--arch |
DPU architecture configuration file for VAI_C compiler in JSON format. It contains the dedicated options for the DPU. We used the configuration file for the ZCU102 board, located at /opt/vitis_ai/compiler/arch/dpuv2/ZCU102/ZCU102.json |
--frozen_pb |
Path of TensorFlow frozen protobuf file. It is the deploy_model.pg file we generated during the quntization flow. |
--output_dir |
Path of output directory after compilation process. We choose ‘coutput/output_ZCU102/’ |
--net_name |
Name of DPU kernel for network model after compiled, its skinl |
For convenience we create a bash script dnnc_skinl.sh with the compilation function call. We update the script with all necessary parameters and execute it within the Docker container under conda environment vitis-ai-tensorflow
$ ./dnnc_skinl.sh
Depending on the node operators in the input network the compilation flow generates two categories of kernels. In our case, the skinl_0 is the accelerator kernel which is to be executed by the DPU in the HW. This kernel is composed within the generated object file dpu_skinl_0.elf in the coutput/output_ZCU102 folder. And skinl_1 is the softmax kernel created to be executed by the cpu. However, in this application we gather the input and output dimensions from the skinl_1 kernel parameters and use them to create a Python function CPUCalculateSoftmax(..) which does the softmax calculation in the host cpu. More details on this are covered in the evaluation section below. Compilation also generates a meta.json file in the same folder. This file contains runtime library paths. We copy the /coutput/ folder using scp to the ZCU102 board. A portion of the output of the compilation flow is shown below:
Kernel topology "skinl_kernel_graph.jpg" for network "skinl"
kernel list info for network "skinl"
Kernel ID : Name
0 : skinl_0
1 : skinl_1
4.0 Model Evaluation on ZCU102
inx’s ZCU102 is an excellent platform for evaluating accelerated machine learning applications targeted for edge deployment. This board features a Zynq UltraScale+ device, DDR4 memory, a rich set of peripherals and interfaces and expansion capability.[8]
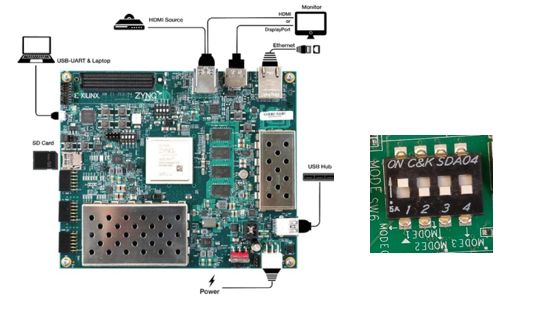

We configure the ZCU102 board so that it boots from an SD card, and connect it to the network via its ethernet port. For simplicity and ease of application evaluation we create a board image with a pre-installed Jupyter notebook. The Jupyter notebook launches automatically at the end of platform boot. We access the Jupyter notebook on a browser at http://board_ip_address:8888. We gather the board_ip_address by accessing it via jtag-over-usb from a laptop. For more details on the configuration options, settings and accessing it via jtag and SSH refer to the user guide UG1414.
Creating the application:
Once the board is up and running we create the lesion detection application in the Jupyter notebook environment. Vitis AI applications can be created using C++ or Python APIs. All required API libraries are already installed in the board image. This image is available for download from the GitHub link. We create the application using Python APIs.
The first step of creating a Vitis AI application in Python is to transform the DPU object file, dpu_skinl_0.elf, we generated in the network compilation step above into a shared library. We achieve this by using the Arm GCC toolchain. Refer to the Jupyter notebook skinl_app.ipynb. The naming format of the shared library must be “libdpumodelModelName.so”, in our case its libdpumodelskinl.so. We place the shared library in the /dpuv2_rundir/ subdirectory which is in the same path level of the skinl_app.ipynb script. The command used is as follows:
# Transform the dpu_skinl_0.elf into shard library libdpumodelskinl.so
!/usr/bin/aarch64-xilinx-linux-gcc -fPIC -shared ../coutput/output_ZCU102/dpu_skinl_0.elf -o libdpumodelskinl.so
Vitis AI comes with a ‘Runner Class’ which provides the abstraction for FPGA initialization and methods for scheduling compute tasks to the DPU. We instantiate the Vitis Runner object with the syntax shown below; the /vitis_rundir/ is the path to the folder containing libdpumodelskinl.so and meta.json files.
# Instantiate the runner for the DPU
dpu = runner.Runner(path+'/dpuv2_rundir/' )
Now we place the input image data in the memory in a format such that DPU can read and process. To do that we gather the input and output tensors of the DPU using the following API calls. The outputs are in Python list format.
# get the input and output tensors
inputTensors = dpu.get_input_tensors()
outputTensors = dpu.get_output_tensors()
We estimate the input and output dimensions from the input and output Tensors lists as follows:
shapeIn = (batchSize,) + tuple([inputTensors[0].dims[i] for i in range(inputTensors[0].ndims)][1:])
outputHeight = outputTensors[0].dims[2]
outputWidth = outputTensors[0].dims[3]
outputChannel = outputTensors[0].dims[1]
We use these dimensions to create input and output data buffers in the memory:
outputData.append(np.empty((batchSize,outputSize), dtype = np.float32, order = 'C'))
inputData.append(np.empty((shapeIn), dtype = np.float32, order = 'C'))
We then populate the input buffer with the lesion images we want to do detection as follows:
#place input images in to the input buffer
k = steps * batchSize
for j in range(batchSize):
imageRun = inputData[0]
imageRun[j,...] =img[j+k].reshape(inputTensors[0].dims[1],inputTensors[0].dims[2],inputTensors[0].dims[3])
Finally, we schedule the compute task to the DPU with the location of the input buffer and the output buffer, where it dumps the computed outputs. Once DPU execution completes the results are available in the outputData buffer.
dpu.execute_async(inputData,outputData)
Finally, we execute the Softmax computation in the CPU by calling CPUCalcSoftmax(..) with the outputData. The CPUCalcSoftmax(..) is a Python function defined in the skil_aux.py file.
CPUCalcSoftmax(outputData[0][j], outputSize)
The output of the Softmax function is a list of detected classes of the batch of lesion images we fed via inputData buffer. In the Jupyter notebook skinl_app.ipnb we used 600 images and demonstrated the estimation of the model accuracy and the average frame/sec ( FPS) of the application.
Screen captures from Jupyter notebook running on zcu102 platform:
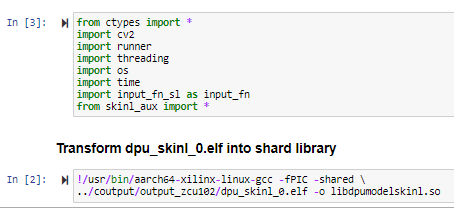
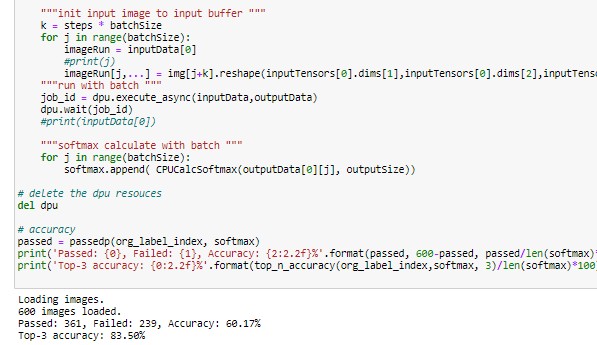
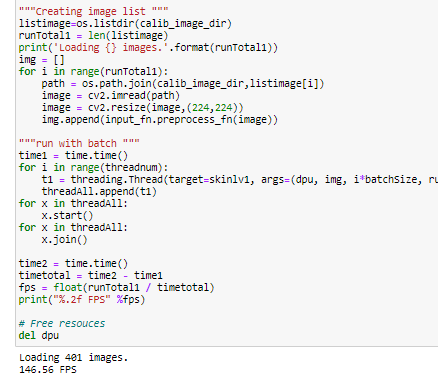
The GitHub link in the references section contains the following items. These are available for download,
1) The Vitis AI tools docker Jupyter notebook dataaug.ipynb, training.ipynb, and related files and libraries
2) The SD image for ZCU102 board with skinl_app.ipynb and related files and necessary libraries.
Acknowledgments
I would like to thank Pat McGuire for the review, edit and thoughtful feedback, and Harvard Dataverse for sharing the Skin Lesion dataset.
Reference
Files located https://github.com/Xilinx/Vitis-AI-Tutorials/tree/Medical-AI-Acceleration
[1] Tschandl, Philipp, 2018, "The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions", https://doi.org/10.7910/DVN/DBW86T, Harvard Dataverse, V1, UNF:6:IQTf5Cb+3EzwZ95U5r0hnQ
[2] ImageNet, An image database organized according to the WordNet hierarchy, http://www.image-net.org/
[3] Xilinx, PG338, DPU for Convolutional Neural Network v3.0 /content/xilinx/en/support/documentation/ip_documentation/dpu/v3_0/pg338-dpu.pdf
[4] Scientific Data, 14 August 2018, Preprocessed and processed image: https://www.nature.com/articles/sdata2018161
[5] Xilinx, UG1414 Vitis AI User Guide, Version 1.0, 10 December 2019, /content/xilinx/en/support/documentation/sw_manuals/vitis_ai/1_0/ug1414-vitis-ai.pdf
[6] Xilinx, The Vitis™ AI software Docker image on Docker hub. https://hub.Docker.com/r/xilinx/vitis-ai/tags
[7] docker, Instructions on installing Docker and add user to the Docker group. https://docs.Docker.com/install/linux/Docker-ce/ubuntu/
[8] Xilinx, ZCU102 product page on the Xilinx website: /content/xilinx/en/products/boards-and-kits/ek-u1-ZCU102-g.html
[9] Google, Image Classification Transfer Learning with Inception v3, https://codelabs.developers.google.com/codelabs/cpb102-txf-learning/index.html#1
[10] Xilinx, Alimul Hoque, https://github.com/Xilinx/Vitis-AI-Tutorials/tree/Medical-AI-Acceleration

About Alimul Hoque
Alimul Hoque is an acceleration solution architect. He joined AMD 3 years ago and is focused on enabling customers creating acceleration solutions for datacenter and edge deployment. Prior to joining AMD Alimul worked for Intel corporation for 10+ years in platform design and datacenter architecture. He holds a BSEE and a MSSE from Northeastern University. He is happily married to Moushumi Nazneen; loves traveling, gardening and watching movies.